Single Cell Proteomic and Transcriptomic Analyses with Single Molecule Sensitivity
Quantifying E. coli Proteome and Transcriptome with Single-Molecule Sensitivity in Single Cells
Gene expression is often stochastic, because gene regulation takes place at a single DNA locus within a cell. Thus, protein and messenger RNA (mRNA) copy numbers vary from cell to cell in isogenic bacterial populations. However, these molecules often exist in low copy numbers and are difficult to detect in single cells, making single molecule detection important.
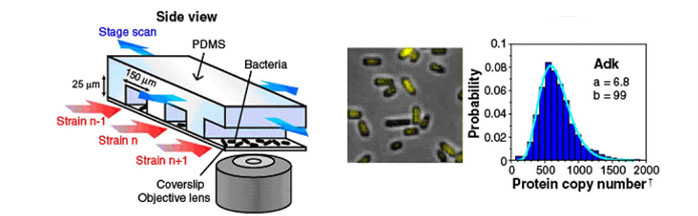
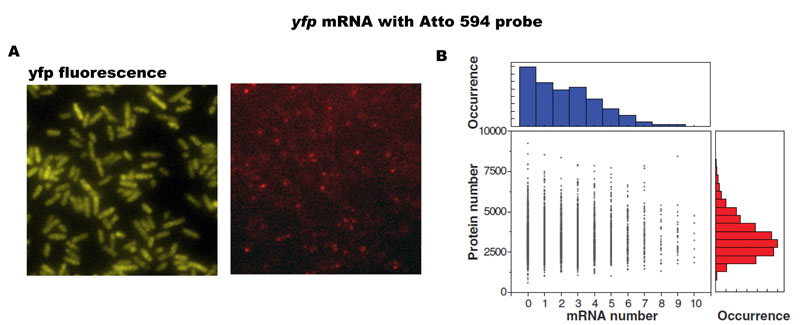
The simultaneous profiling of mRNA and protein revealed that the mRNA and protein copy numbers of a single cell for any given gene are uncorrelated (Figure 2); that is, a cell that has more mRNA molecules than average does not necessarily have more proteins. This perhaps counter-intuitive result can be explained by the fact that mRNA has a much shorter lifetime than protein in bacteria. A mammalian cell, by contrast, has comparable mRNA and protein lifetimes, and hence is expected to have more correlated mRNA and protein levels than a bacterial cell. Taken together, a quantitative and integral account of a single-cell gene expression profile is emerging.
References:
[1] Taniguchi, Y., Choi, P.J., Li, G.-W., Chen, H., Babu, M., Hearn, J., Emili, A., Xie, X.S. (2010), “Quantifying E. coli Proteome and Transcriptome with Single-Molecule Sensitivity in Single Cells”, Science 329: 533-538
[2] Friedman, N., Cai, L., Xie, X.S. (2006), “Linking Stochastic Dynamics to Population Distribution: An Analytical Framework of Gene Expression”, Phys. Rev. Lett. 97: 168302