Single Cell Sequencing by LIANTI
Single-cell genomics is important for biology and medicine. However, previous whole-genome amplification (WGA) methods have limited capability to detect copy-number variations (CNVs) and single-nucleotide variations (SNVs). This can be attributed to the low amplification evenness and fidelity by exponential amplification (Figure 1).
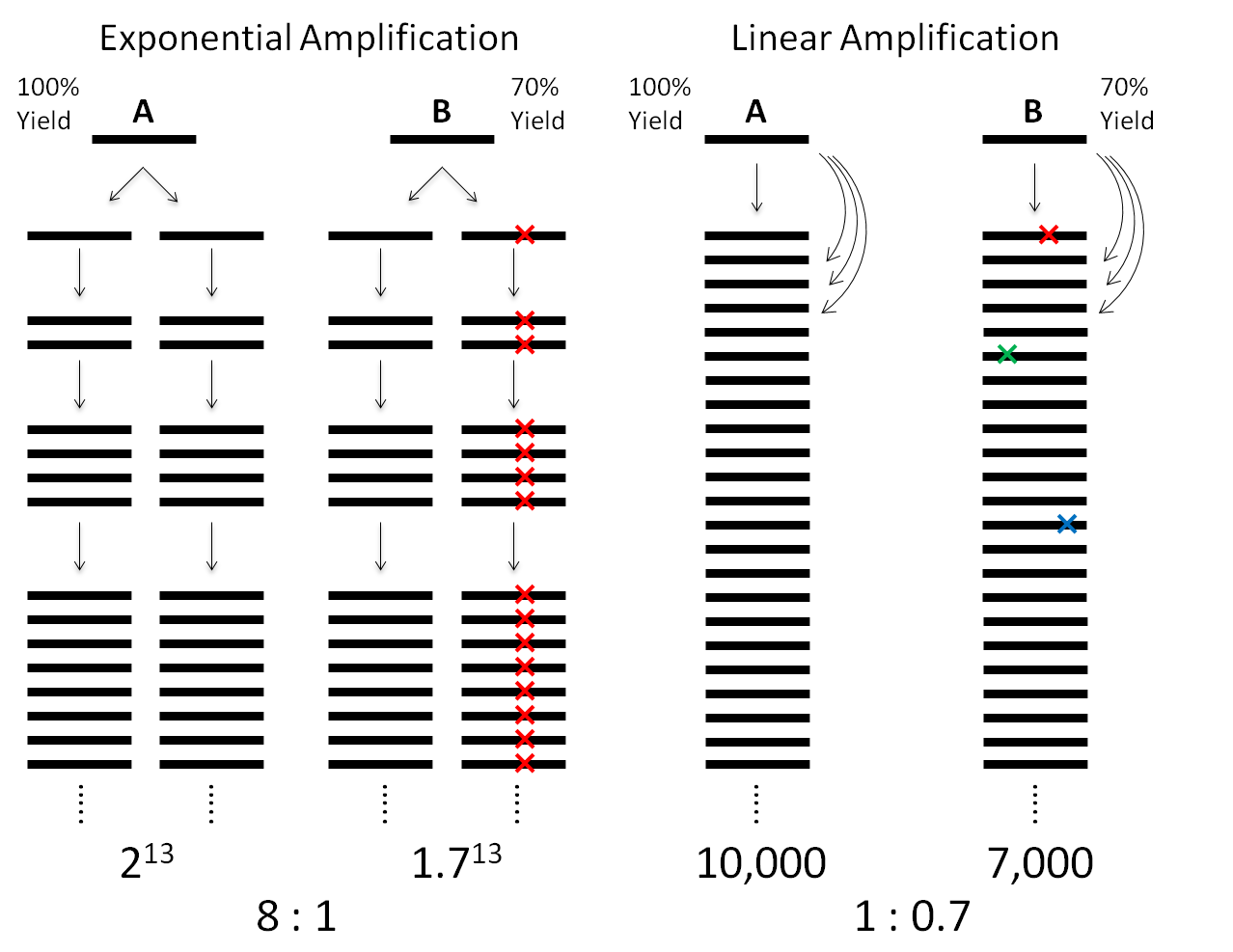
To overcome the limitations of exponential amplification, we have developed a new single-cell WGA method, Linear Amplification via Transposon Insertion (LIANTI). LIANTI takes advantage of Tn5 transposition and T7 in vitro transcription to linearly amplify genomic DNA fragments from a single human cell (Figure 2).
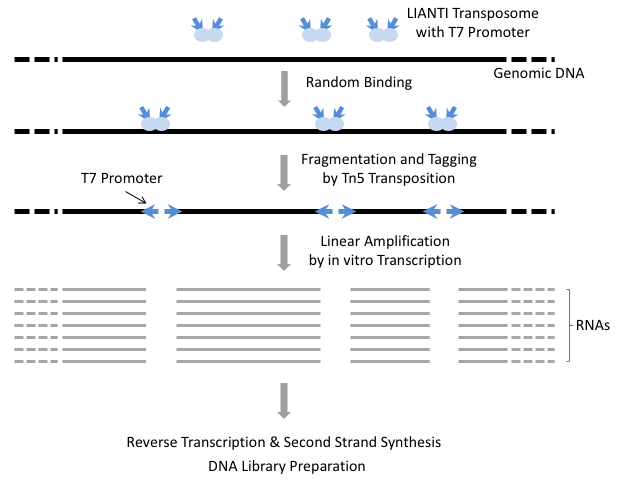
LIANTI exhibits the highest amplification uniformity compared to the other methods (Figure 3A, 3B). Together with its unique capability of digital counting to infer fragment numbers based on the mapping coordinates of each amplicon, LIANTI allows accurate detection of single-cell micro-CNVs with kilobase resolution (Figure 3C, 3D).
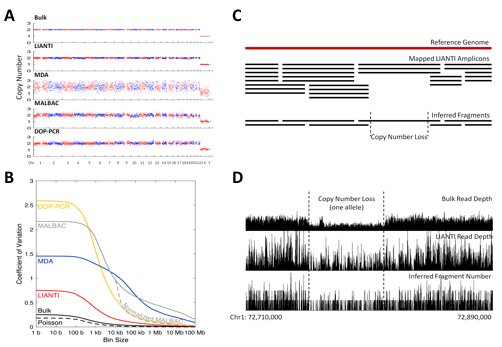
LIANTI also achieves the highest amplification fidelity for accurate single-cell SNV detection (Figure 4A). Furthermore, we show that the predominant cytosine-to-thymine mutations observed in single-cell genomics are often SNV false positives arising from the artifact of cytosine deamination upon cell lysis (Figure 4B).
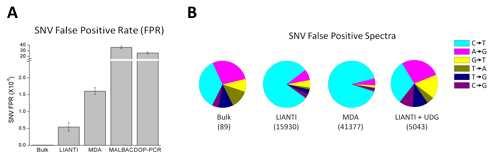
Using LIANTI, we directly observed the stochastic firing of DNA replication origins, which differs from cell to cell. We also determined the spectrum and genome-wide distribution of ultraviolet-induced SNVs in a single human cell. In addition, the high precision of micro-CNVs detection and the ability to call individual SNVs in a single cell will allow better genetic screening in reproductive medicine and provide valuable information about how genome variation takes place in cancer and other diseases.
References:
- Chen, Chongyi; Xing, Dong; Tan, Longzhi; Li, Heng; Zhou, Guangyu; Huang, Lei; Xie, X. Sunney “Single-cell whole-genome analyses by Linear Amplification via Transposon Insertion (LIANTI),” Science 356, 189-194. DOI:10.1126/science.aak9787 (2017)