Non-Specific DNA-Protein Interactions
DNA repair proteins slide in contact with DNA to find defect bases
Many DNA-binding proteins need to locate target sites in DNA very quickly. These proteins face a difficult needle-in-a-haystack problem: the rare target sites are hidden among billions of nonspecific sites in a large DNA genome. These proteins are thought to solve the search problem through a combination of “hopping” among different DNA segments and “sliding” along DNA segments. While there exists evidence for diffusion of proteins along DNA [1], it has not been clear if this took place exclusively by hopping or whether the proteins can slide in contact with the DNA.

We have combined the DNA flow-stretching assay with single-molecule fluorescence detection to monitor single fluorescently-labeled DNA-binding proteins interacting with stretched DNA molecules in real-time [2]. For these measurements, we do not add a bead to the free end of the DNA. Using wide-field total internal reflection fluorescence microscopy, we can image the DNA-bound protein molecules while minimizing the fluorescence background from protein molecules in solution.
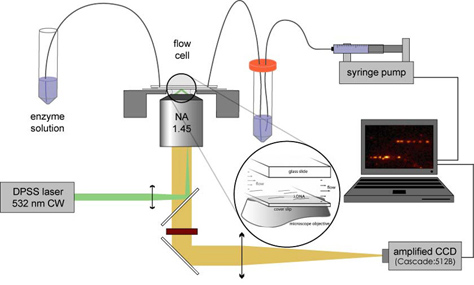
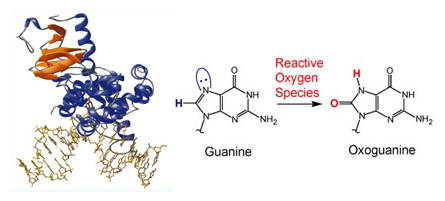
We label hOgg1 with the fluorescent dye Cy3B at the protein’s C-terminus. When a small amount of the labeled protein is added to the flow solution, we observe that the DNA repair enzymes attach to and move along the flow-stretched DNA strands. Over the course of a few minutes we see hundreds to thousands of protein molecules attach to, move along, and dissociate from each DNA strand in the field of view. By determining the positions of the molecules in each frame of a recorded movie, we can trace out the path each protein molecule took as it moved along DNA searching for OxoG.
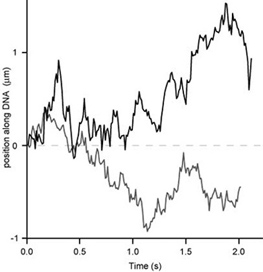
Using many such molecular trajectories, we can construct population-averaged mean-square displacement plots to accurately measure the protein’s diffusion constant under a given buffer condition.
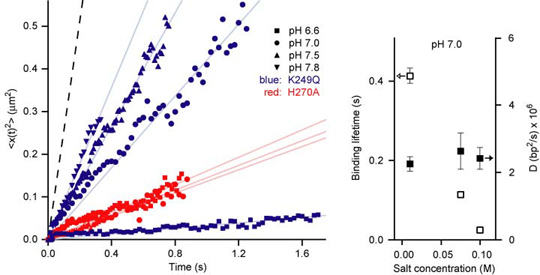
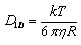
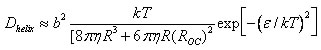
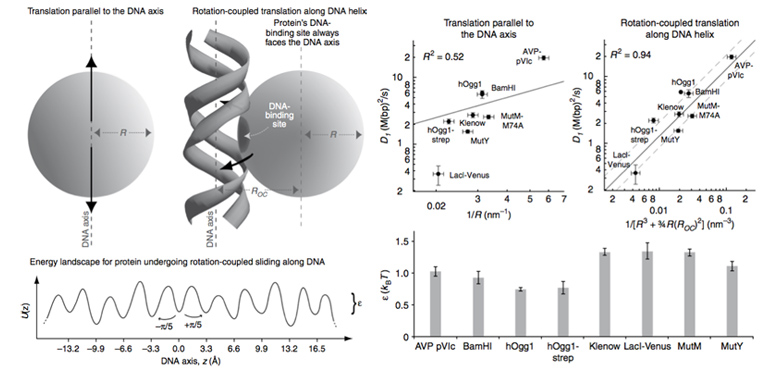
As shown in the above figure, the size dependence of the measured apparent 1D diffusion constants indicates that DNA-binding proteins undergo rotation-coupled sliding along the DNA helix rather than sheer translation [4]. The fact that eight proteins of varying size are consistent with the theory of spinning along the DNA helix in a rugged free-energy landscape [6] indicates that rotation-coupled sliding is a general phenomenon. The average free-energy barrier ε is 1.1 ± 0.2 kBT. Such small barriers facilitate rapid search for binding sites.
References
[1] Berg, O.G.; Winter, R.B; von Hippel, P.H. “Diffusion-Driven Mechanisms of Protein Translocation on Nucleic Acids. 1. Models and Theory?” Biochemistry 20, 6929 (1981).
[2] Blainey, P. C.; van Oijen, A. M.; Banerjee, A.; Verdine, G. L.; Xie, X. S. “A base-excision DNA-repair protein finds intrahelical lesion bases by fast sliding in contact with DNA,” Proc. Natl. Aca. Sci., 103, 5752-5757 (2006).
[3] Bruner, S. D.; Norman, D. P. G.; Verdine, G. L. “Structural Basis for Recognition and Repair of the Endogenous Mutagen 8-oxoguanine in DNA,” Nature 403, 859-866 (2000).
[4] Blainey, P. C.; L.Guobin; Kou, S.C.; Mangel, W. F.; Verdine, G. L.; Bagchi, B.; Xie, X. S. “Nonspecifically bound proteins spin while diffusing along DNA” Nature Structural & Molecular Biology, 16, 1224-1229 (2009).
[5] Schurr, J.M. “The one-dimensional diffusion coefficient of proteins absorbed on DNA. Hydrodynamic considerations,”Biophys. Chem. 9, 413–414 (1979).
[6] Bagchi, B.; Blainey, P. C.; Xie, X. S. “Diffusion Constant of a Nonspecifically Bound Protein Undergoing Curvilinear Motion along DNA,” J.Phys.Chem.B, 112, 6282-6284 (2008).