Single-Molecule Imaging inside a Live Mammalian Cell Nucleus
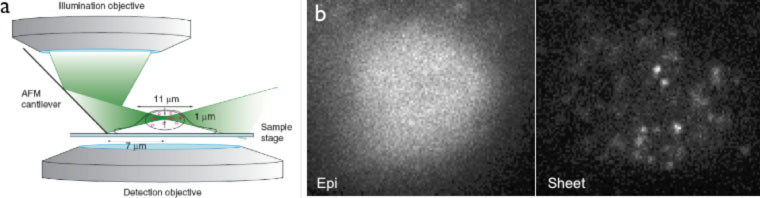
Tracking single molecules in living cells provides a direct way to probe the kinetics of their interactions with other cellular components, and is particularly useful to characterize unsynchronized dynamic events. Despite previous successes in single-molecule imaging in live bacterial cells, imaging single fluorescent proteins remains challenging in the nuclei of living mammalian cells, due to their larger size and higher fluorescence background. We developed a novel fluorescence microscopy technique that combines selective plane illumination with a vertical arrangement of illumination and detection objectives. In this new geometry (Fig. 1), a disposable mirror reflects the light sheet into a horizontal plane close to the sample surface, thus allowing horizontal sectioning of the cells and achieving single fluorescent protein imaging in live mammalian cells. We name this technique reflected light sheet microscopy (RLSM).
Our technique possesses the advantage of superior signal-to-background ratio, fast image acquisition speed and millisecond time resolution, low photobleaching rate and reduced phototoxicity, as well as easy optical sectioning capability. The reflected light sheet configuration also allows individual, normal-sized adherent cells to be imaged, as compared to other selective plane illumination methods that could only be applied to embryos or cellular spheroids much larger in size. Lastly, the single-molecule tracking capability of RLSM allows us to directly monitor in vivo protein dynamics without the need for additional calibrations or corrections that are commonly associated with other techniques such as FRAP and FCS.
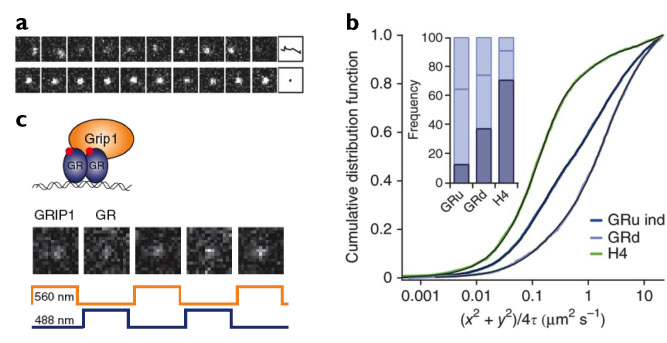
We demonstrate the potential of our new microscopy method by directly monitoring the binding properties of fluorescently labeled glucocorticoid receptors (GR) to DNA (Fig. 2a and b). GR is a transcription factor that localizes mostly to the cytoplasm in the absence of hormone, but forms homodimers and translocates into the nucleus upon binding to hormone. Previous studies have shown that dimeric GR binds directly to DNA at regulatory sequences, while the monomer can be indirectly recruited to DNA by other DNA-bound protein complexes. The mode of DNA interaction defines whether the target gene is activated or repressed. Using RLSM, we measured the DNA-bound fraction of GR and determined the residence times on DNA of the various oligomerization states of GR, thereby permitting us to resolve the transcription factor’s different modes of DNA binding. We also demonstrated the capability of RLSM for two-color single-molecule imaging (Fig. 2c) by directly observing the spatio-temporal co-localization of GR and its coactivator GRIP1. The imaging technique we have developed could be generally applied to study the dynamics of a variety of key biomolecular players inside the living mammalian nucleus.
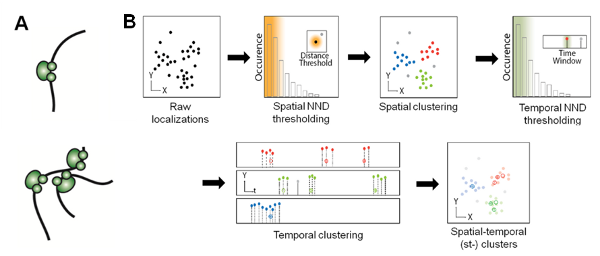
In addition, we combined RLS illumination with super-resolution microscopy (SRM), and applied RLS-SRM to image the spatial organization of RNAP II molecules inside the mammalian nucleus, which has long been proposed to heterogeneously occur in discrete foci termed ‘transcription factories’ (Fig. 3A). Leveraging on the blinking photophysics of rhodamine-based dyes such as TMR, we also developed a density-based clustering algorithm that pools multiple localizations in super-resolution images based on their spatial and temporal proximity, so as to be able to accurately count the copy number of RNAP II molecules in transcription foci (Fig. 3B).
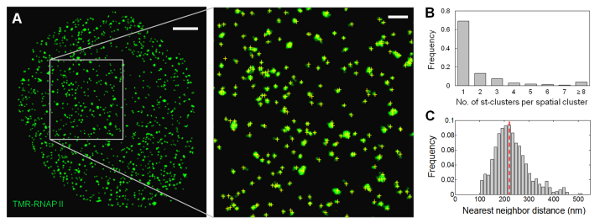
Using RLS-SRM, we quantified the global extent of RNAP II clustering inside the mammalian nucleus. We found that the majority (< 70%) of the transcription foci originate from single RNAP II molecules, and no significant clustering between RNAP II molecules is detected within the length scale of the reported diameter of ‘transcription factories’ (Fig. 4). Colocalization measurements of RNAP II molecules equally labeled by two spectrally distinct dyes confirmed the primarily unclustered distribution, arguing against a prevalent existence of ‘transcription factories’ in the mammalian nucleus as previously proposed. Our study not only provides clear and convincing answers to this long-standing controversy, but also paves way for quantitative mapping and stoichiometric characterization of other biomolecular species deep inside mammalian cells.
References:
Gebhardt, J. Christof M.; Suter, David M.; Roy, Rahul; Zhao, Ziqing W.; Chapman, Alec R.; Basu, Srinjan; Maniatis, Tom; Xie, X. Sunney. “Single-Molecule Imaging of Transcription Factor Binding to DNA in Live Mammalian Cells,” Nat Methods 10, 421-426 DOI:10.1038/NMETH.2411 (2013)
Zhao, Ziqing W.; Roy, Rahul; Gebhardt, J. Christof M.; Suter, David M.; Chapman, Alec R.; Xie, X. Sunney. “Spatial organization of RNA polymerase II inside a mammalian cell nucleus revealed by reflected light-sheet superresolution microscopy,” PNAS 111, 681-686 DOI:10.1073/pnas.1318496111 (2014)